Show/hide the code
# create function for creating an outcome table
make_outcome_block <- function(data, outcome_var, outcome_label) {
# convert outcome_var to a symbol so we can use !!
outcome_var <- rlang::ensym(outcome_var)
# summarise outcome by time and group (numeric means/sds)
long <- data |>
select(group, event, value = !!outcome_var) |>
mutate(
time = case_when(
event == 0 ~ "Baseline",
event == 12 ~ "12-weeks",
event == 26 ~ "26-weeks",
TRUE ~ as.character(event)
),
time = factor(
time,
levels = c("Baseline", "12-weeks", "26-weeks"),
ordered = TRUE
)
) |>
filter(!is.na(value)) |>
group_by(group, time) |>
summarise(
mean = mean(value, na.rm = TRUE),
sd = sd(value, na.rm = TRUE),
.groups = "drop"
)
# numeric means wide by time (for deltas)
mean_wide <- long |>
select(group, time, mean) |>
tidyr::pivot_wider(
id_cols = group,
names_from = time,
values_from = mean,
names_glue = "{time}_mean"
)
# ensure all time columns exist, even if NA
for (tn in c("Baseline_mean", "12-weeks_mean", "26-weeks_mean")) {
if (!tn %in% names(mean_wide)) mean_wide[[tn]] <- NA_real_
}
mean_wide <- mean_wide |>
mutate(
delta = `26-weeks_mean` - `Baseline_mean`
)
# formatted mean (SD) wide by time
cell_wide <- long |>
mutate(
mean = round(mean, 1),
sd = round(sd, 1),
cell = sprintf("%.1f (%.1f)", mean, sd)
) |>
select(group, time, cell) |>
tidyr::pivot_wider(
id_cols = group,
names_from = time,
values_from = cell,
names_glue = "{time}_cell"
)
# ensure all formatted time columns exist
for (tn in c("Baseline_cell", "12-weeks_cell", "26-weeks_cell")) {
if (!tn %in% names(cell_wide)) cell_wide[[tn]] <- NA_character_
}
# merge numeric and formatted,
full <- mean_wide |>
left_join(cell_wide, by = "group") |>
mutate(
delta_str = ifelse(
is.na(delta),
NA_character_,
sprintf("%.1f", round(delta, 1))
)
) |>
transmute(
Label = as.character(group),
Baseline = `Baseline_cell`,
`12-weeks` = `12-weeks_cell`,
`26-weeks` = `26-weeks_cell`,
`Δ 26w – baseline` = delta_str
)
# add outcome header row on top
header_row <- full[0, , drop = FALSE]
header_row[1, ] <- NA_character_
header_row$Label[1] <- outcome_label
tab <- bind_rows(header_row, full)
# force all columns to character to bind_rows across outcomes
tab[] <- lapply(tab, as.character)
tab
}
# apply function to outcomes
# create table
out_table <- bind_rows(
make_outcome_block(df, gds_score_new, "Depression (GDS-SF)"),
make_outcome_block(df, smast_score_new, "Alcohol use (SMAST-G)"),
make_outcome_block(df, sidas_score, "Suicide Risk (SIDAS)"),
make_outcome_block(df, lonely_total, "Loneliness (UCLA Loneliness Scale)")
)
out_table <- out_table |>
mutate(across(everything(), ~ ifelse(is.na(.), "", .)))
# create list of outcome labels
outcome_labels <- c(
"Depression (GDS-SF)",
"Alcohol use (SMAST-G)",
"Suicide Risk (SIDAS)",
"Loneliness (UCLA Loneliness Scale)"
)
out_table <- out_table |>
flextable() |>
my_flextable() |>
padding(i=c(2:3, 5:6, 8:9, 11:12), j=1, padding.left=20) |>
set_header_labels(Label = "Outcome") |>
align(j = c(2:5), part = "all", align = "center")
out_tableOutcome | Baseline | 12-weeks | 26-weeks | Δ 26w – baseline |
|---|---|---|---|---|
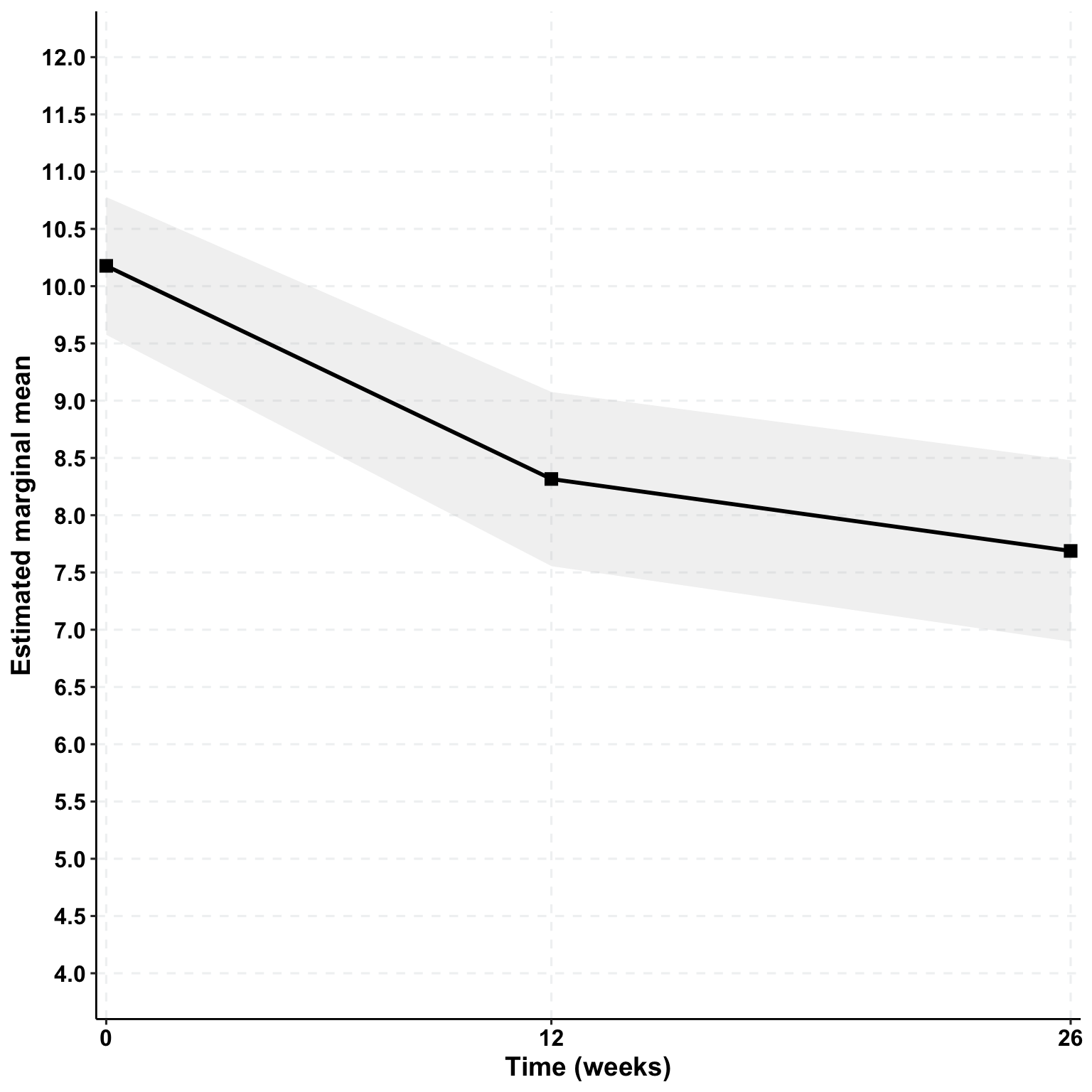
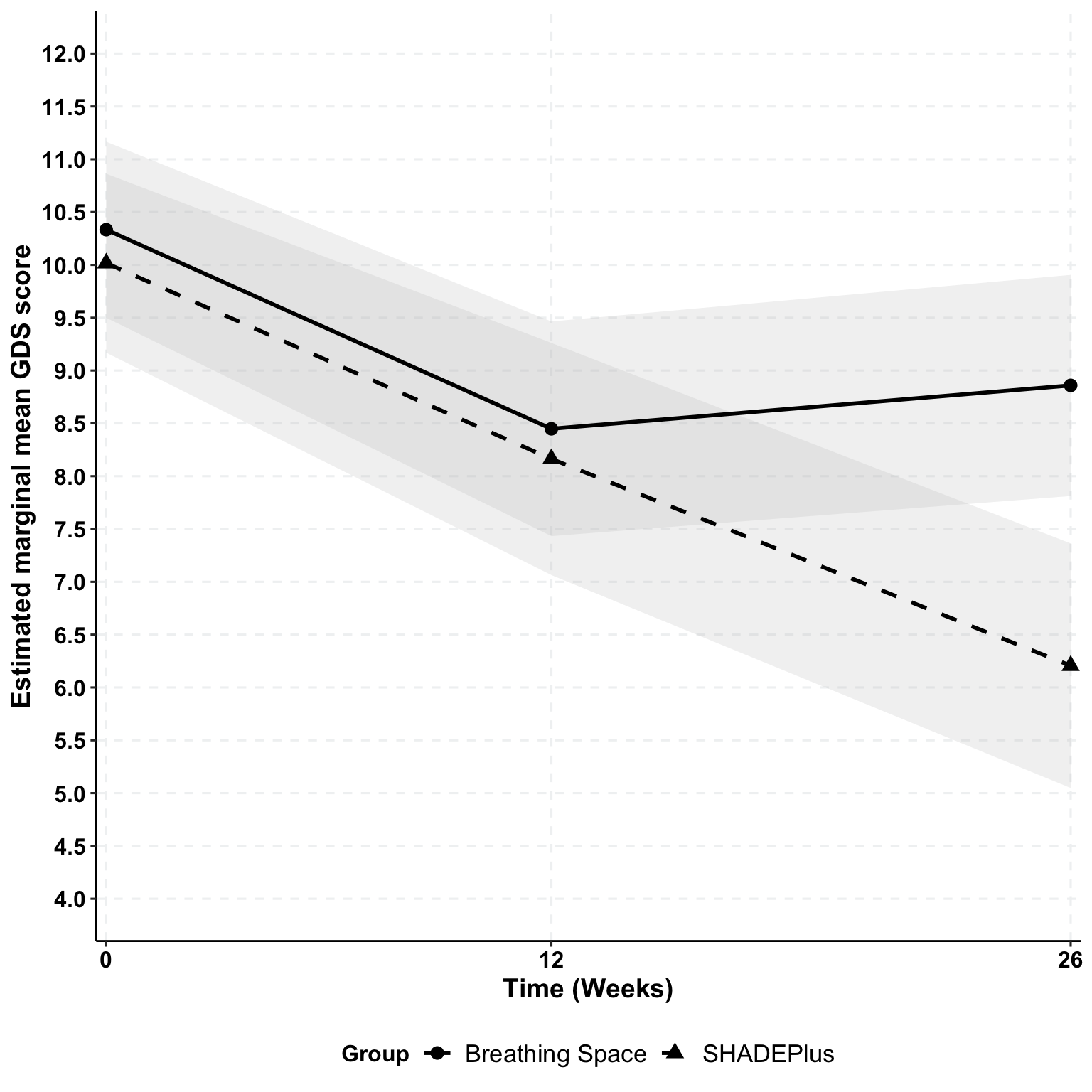
Depression (GDS-SF) | ||||
Breathing Space | 10.3 (2.9) | 8.3 (3.7) | 8.8 (4.1) | -1.5 |
SHADEPlus | 10.0 (2.9) | 8.1 (3.8) | 6.1 (4.2) | -3.9 |
Alcohol use (SMAST-G) | ||||
Breathing Space | 6.4 (2.2) | 5.6 (2.4) | 5.7 (2.4) | -0.7 |
SHADEPlus | 6.2 (2.1) | 5.1 (2.9) | 4.1 (2.7) | -2.1 |
Suicide Risk (SIDAS) | ||||
Breathing Space | 12.3 (8.2) | 10.1 (8.0) | 9.8 (10.7) | -2.6 |
SHADEPlus | 13.2 (8.1) | 12.1 (8.5) | 9.5 (12.0) | -3.7 |
Loneliness (UCLA Loneliness Scale) | ||||
Breathing Space | 7.5 (1.6) | 7.2 (1.4) | 7.7 (1.6) | 0.2 |
SHADEPlus | 7.3 (1.7) | 6.6 (1.9) | 6.6 (2.2) | -0.7 |